9. beta_PCA.py¶
9.1. Description¶
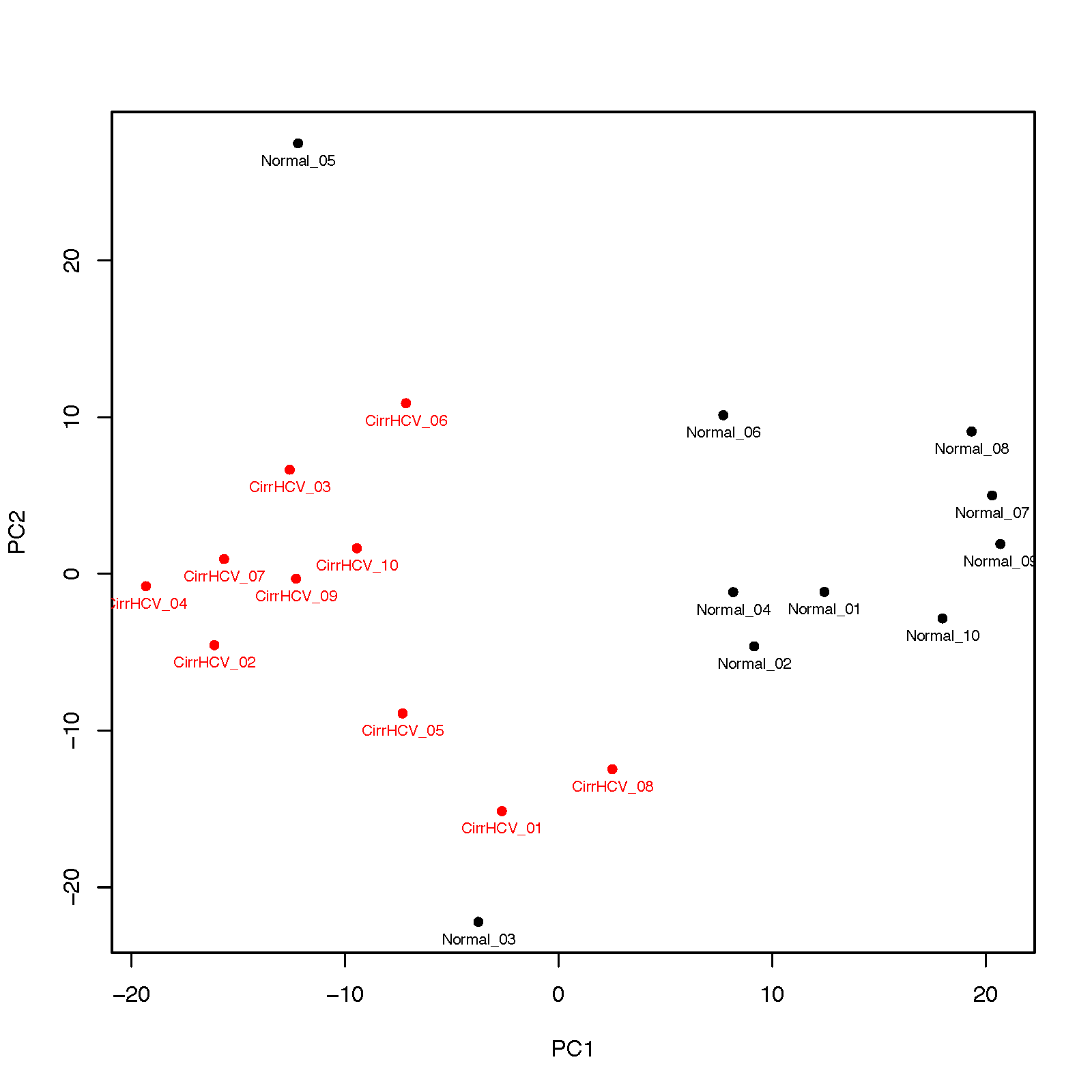
This program performs PCA (principal component analysis) for samples.
Example of input data file
ID Sample_01 Sample_02 Sample_03 Sample_04
cg_001 0.831035 0.878022 0.794427 0.880911
cg_002 0.249544 0.209949 0.234294 0.236680
cg_003 0.845065 0.843957 0.840184 0.824286
...
Example of input group file
Sample,Group
Sample_01,normal
Sample_02,normal
Sample_03,tumor
Sample_04,tumo
...
Notes
Rows with missing values will be removed
Beta values will be standardized into z scores
Only the first two components will be visualized
Variance% explained by each component will be printed to screen
9.2. Options¶
- --version
show program’s version number and exit
- -h, --help
show this help message and exit
- -i INPUT_FILE, --input_file=INPUT_FILE
Tab-separated data frame file containing beta values with the 1st row containing sample IDs and the 1st column containing CpG IDs.
- -g GROUP_FILE, --group=GROUP_FILE
Comma-separated group file defining the biological groups of each sample. Different groups will be colored differently in the PCA plot.
- -n N_COMPONENTS, --ncomponent=N_COMPONENTS
Number of components. default=2
- -o OUT_FILE, --output=OUT_FILE
The prefix of the output file.
9.3. Input files (examples)¶
9.4. Command¶
$beta_PCA.py -i cirrHCV_vs_normal.data.tsv -g cirrHCV_vs_normal.grp.csv -o HCV_vs_normal